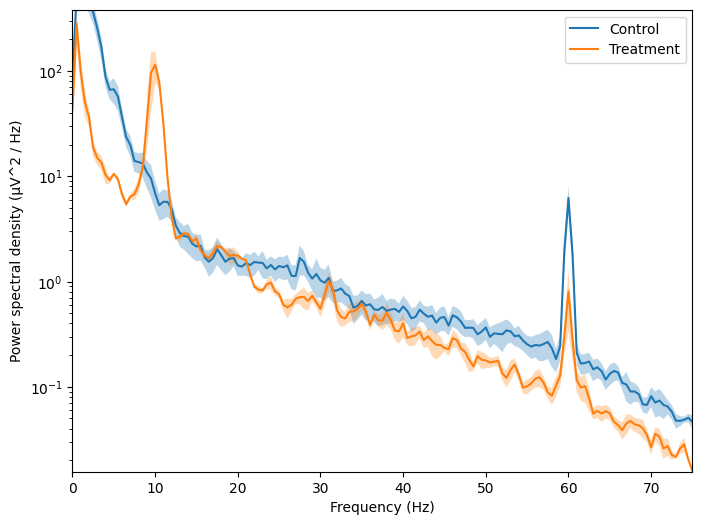
Compute average PSD#
This notebook builds on the EEG analysis in lab to compute an average PSD for different conditions.
Step 1#
Export all of the LabChart Text Files (.txt) for your conditions separately. Then, run the cell below which will create as many folders for you as you have conditions.
# Rename the conditions here!
conditions = ['Control', 'Treatment']
for condition in conditions:
!mkdir -p {condition}
print("Directories created for conditions: " + ", ".join(conditions))
Directories created for conditions: Control, Treatment
Step 2#
Upload your exported .txt files into their respective folders.
import numpy as np
import matplotlib.pyplot as plt
from scipy import signal
import glob
filenames = {}
for condition in conditions:
print(f"Files for {condition}:")
filenames[condition] = glob.glob(f'{condition}/*.txt')
print(filenames[condition])
Files for Control:
['Control/eyes open .txt', 'Control/EEG_Trial1_EyesOpen.txt', 'Control/EEG Eyes Open.txt']
Files for Treatment:
['Treatment/eyes closed.txt', 'Treatment/EEG_Trial1_EyesClosed.txt', 'Treatment/EEG Eyes Closed.txt']
Step 3#
Run the cell below to define several helper functions. You should also change the sampling frequency if you changed it for some reason.
# Change this if it was different for your experiment!
sampling_freq = 400
nperseg = 800 # Window length in samples (2 seconds at 400 Hz)
def generate_avg_psd(filenames):
if len(filenames) == 0:
return np.array([]), np.array([]), np.array([])
psd_list = []
freqs = None
for file in filenames:
print(file)
columns = ['time', 'recording']
data = np.genfromtxt(file, dtype=float, usecols=(0,1), skip_header=6, delimiter='\t', names=columns, encoding='unicode_escape')
recording = data['recording']
# Interpolate NaN values
nans = np.isnan(recording)
x = np.arange(len(recording))
recording[nans] = np.interp(x[nans], x[~nans], recording[~nans])
freqs, psd_single = signal.welch(recording, sampling_freq, nperseg=nperseg)
psd_list.append(psd_single)
# Interpolate all PSDs to same length
max_len = max(len(p) for p in psd_list)
psd_matrix = np.zeros((max_len, len(filenames)))
for i, psd in enumerate(psd_list):
if len(psd) < max_len:
psd = np.interp(np.linspace(0, 1, max_len), np.linspace(0, 1, len(psd)), psd)
psd_matrix[:, i] = psd
average_psd = np.mean(psd_matrix, axis=1)
std_psd = np.std(psd_matrix, axis=1) / np.sqrt(len(filenames))
return average_psd, std_psd, freqs
def plot_psd(average_psd, std_psd, freqs, label=None):
plt.semilogy(freqs, average_psd, label=label)
plt.fill_between(freqs, average_psd - std_psd, average_psd + std_psd, alpha=0.3)
plt.xlabel('Frequency (Hz)')
plt.ylabel('Power spectral density (µV^2 / Hz)')
max_freq = 75
plt.xlim([0, max_freq])
visible_mask = freqs <= max_freq
ymin = min((average_psd - std_psd)[visible_mask])
ymax = max((average_psd + std_psd)[visible_mask])
plt.ylim(ymin, ymax)
Step 4#
Run the cell below to use the functions to calculate your average PSD and then plot for all of your conditions.
average_psd = {}
std_psd = {}
plt.figure(figsize=(8, 6))
for condition in conditions:
average_psd[condition], std_psd[condition], freqs = generate_avg_psd(filenames[condition])
plot_psd(average_psd[condition],std_psd[condition],freqs, label=condition)
plt.legend()
plt.show()
Control/eyes open .txt
Control/EEG_Trial1_EyesOpen.txt
Control/EEG Eyes Open.txt
Treatment/eyes closed.txt
Treatment/EEG_Trial1_EyesClosed.txt
Treatment/EEG Eyes Closed.txt